Highlights
- The crystal structures of SecDF were determined at high resolution.
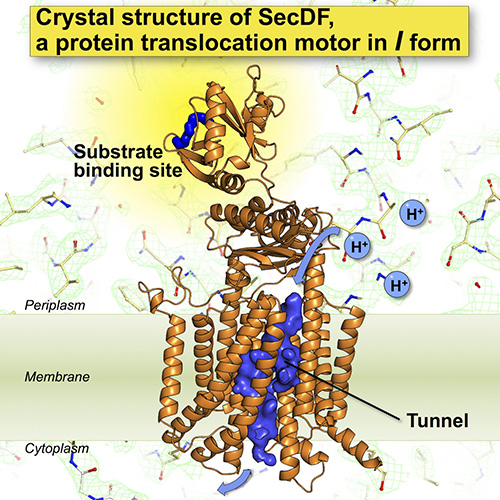
- The structures suggested the presence of a site interacting with protein substrates and a pathway for H+ ion.
- Knowledge of the transmembrane protein export mechanism will provide a structural basis for the development of new type of antibiotics.
Summary
The cell membrane separates the cell's interior from the environment. Extracellular transport of proteins synthesized in the cytoplasm is an essential process for sustaining life. We conduct basic research on the molecular mechanisms of the protein transport.
In bacteria such as Escherichia coli, the membrane-embedded SecDF is involved in protein transport. Specifically, SecDF mediates the translocation of its substrates by using the H+ gradient across the membrane. However, the detailed molecular mechanisms of SecDF remains unclear. In particular, the sites that interact with substrate proteins passing through the membrane and the H+ pathway remained elusive.
Recently, we successfully determined the crystal structure of SecDF at high resolution (2.6 Å). Based on this structural information, we carried out biochemical experiments and computational molecular dynamics simulation. The results of these analyses identified a SecDF's substrate-binding site and a H+ ion pathway. Moreover, we demonstrated that each SecDF domain undergoes dynamic conformational changes, driven by the H+ ion gradient. Based on these findings, we proposed a novel molecular mechanism for SecDF-driven protein translocation across the membrane.
Our findings regarding this essential biological phenomenon will contribute to progress in basic life science. In addition, our findings will provide a theoretical basis for developing new antibiotics that target the bacteria-specific SecDF protein.
For inquiries
Tomoya Tsukazaki
Associate Professor
Membrane Molecular Biology Laboratory
Graduate School of Biological Sciences
Nara Institute of Science and Technology
MAIL: ttsukaza[@]bs.naist.jp
TEL: 0743-72-5551